|
1/13/2024 0 Comments Error bars not at top of graph r  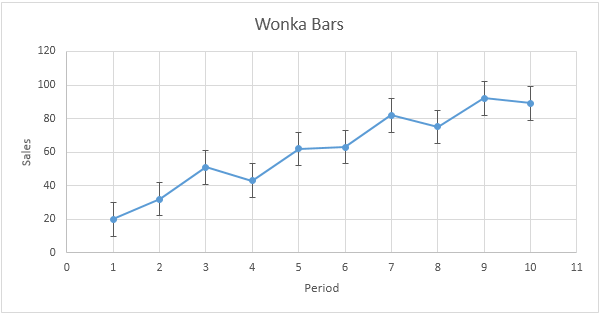
15.4 Calculating the five parameters of all subjects, groups and conditions.15.1 Contrast Sensitivity Function Model.15 Understanding the Contrast Sensitivity Function.14.3 Calculating area and R2 of all subjects, groups and conditions.14 Area under Curve, AULCSF and R2 of the CSF.13.3 Facetting the Contrast Sensitivity Functions.13 Plotting the Contrast Sensitivity Function.12.4.1 Annotation using sm_forest_annot().12.3 A Bland Altman plot - sm_bland_altman().12 Slope Charts, Point plots, Bland-Altman, Forests, Rainclouds, Histograms (Part 2).11.8.1 Plotting individual points with unique colors.11.7.1 Plotting individual points with unique colors.11.6.1 Plotting individual points with unique colors.11.5.4 Correlation plot with both regression and reference lines.11 Themes, Colors, Correlations, Boxplots, Violins and Bars (Part 1).10.7.3 Checking the Assumption for Homogeneity of Variance.10.6.1 Issues with post-hoc power analysis.10.2.1 Shapiro-Wilk Test to test for Normality of Data.9.2 Calculating slopes of all subjects, groups and conditions.8.3.3 Figure 3E (modeling in Matlab and plotting in R).8.3.1 Figure 3B (a best-fit line with points and error bars).8.3 Reproducing figures in the paper (Min et al., 2019).7.2 Plotting the averaged data with error bars.7.1.3 summarise() for grouped summaries.6.2.1 Plotting the forest plot using data of multiple groups.6.1.4 Raincloud plot with data from multiple groups.6.1.3 Changng the theme and orientation.6.1.2 Separating the components and configuration.5.3.4 Point plot with a shadow using data of multiple groups.5.3.3 Point plot with a shadow using data of one group.5.2.4 Slope chart with multiple groups and x levels.5.2.3 Slope chart with multiple x levels.5.2.2 Slope chart with mean plot and error bar.5.1.4 Bar graph with data of multiple groups.5.1.2 Plotting individual points with unique colors.5 Bar Graph, Slope Chart and Point plot.4.3.3 Violin plot with individual points.4.2.6 Double-check if the ‘Day’ column is factor.4.2.5 Displaying characters in a non-alphabetical order.4.1 Upload sample data (csv file) in RStudio.3.3.7 Let’s save the plot as an image in your folder LearnR by using the variable figure1.3.3.6 Different color for each group but with other colors.3.3.4 Plotting the average with standard errors.3.3.2 Reporting statistics from a paired correlation. 
3.3.1 Positive relationship between x- and y-axes.3.3 Improve data visualization using smplot2.3.2.6 How do we draw the best-fit line of the graph?.3.2.4 Different color & shape for each group.3.2.3 Different color of points for each unique group.3 Basics of ggplot2 and Correlation Plot.How can I learn most effectively with the notes?.Let’s make a folder and set it as working directory.Theme_classic() + labs(title= "Cytotoxicity assay", x = "Conc. Stat_summary(fun = mean, position=position_dodge(width=0.95), geom = "line", size = 1) + Like above we will use stat_summary() to plot the error bar first then the draw the lines. a <- ggplot(data, aes(y = Cell, x = Conc., colour = Name, group = Name)) + for x-axis instead of drug types and we will color the lines according to the drug type. Why we are creating another ggplot object because we will use Conc. Now let’s use the same data but draw a line instead. a + stat_summary(fun.data= mean_cl_normal, If we replace mean_sdl with mean_cl_normal we will be asking for 95% confidence intervals. Y = "Cell viability %") + labs(fill='Conc. Theme_classic() + labs(title= "Cytotoxicity assay", x = "", Position=position_dodge(width=0.95),geom="bar") + The highlighted part in red in the command below is telling the function to create the error bar depending on standard deviation. The first stat_summary() function is to plot the error bars and the second stat_summary()function is to plot the bars. This function makes it so easy to visualize the error bars. We will be using stat_summary() function. a <- ggplot(data,aes(x=Name, y=Cell, fill=Conc.)) x-axis will be Name (drug names) and y-axis will be Cell (Cell viability %) and we will use Conc. #load the data from Disk A (specify where is your file, it should be in CSV format) So first we should load ggplot2 library and load our table into RStudio #load ggplot2 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
